Abstract
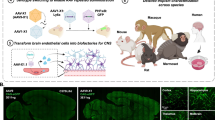
The blood–brain barrier (BBB) restricts the systemic delivery of messenger RNAs (mRNAs) into diseased neurons. Although leucocyte-derived extracellular vesicles (EVs) can cross the BBB at inflammatory sites, it is difficult to efficiently load long mRNAs into the EVs and to enhance their neuronal uptake. Here we show that the packaging of mRNA into leucocyte-derived EVs and the endocytosis of the EVs by neurons can be enhanced by engineering leucocytes to produce EVs that incorporate retrovirus-like mRNA-packaging capsids. We transfected immortalized and primary bone-marrow-derived leucocytes with DNA or RNA encoding the capsid-forming activity-regulated cytoskeleton-associated (Arc) protein as well as capsid-stabilizing Arc 5’-untranslated-region RNA elements. These engineered EVs inherit endothelial adhesion molecules from donor leukocytes, recruit endogenous enveloping proteins to their surface, cross the BBB, and enter the neurons in neuro-inflammatory sites. Produced from self-derived donor leukocytes, the EVs are immunologically inert, and enhanced the neuronal uptake of the packaged mRNA in a mouse model of low-grade chronic neuro-inflammation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The data supporting the results in this study are available within the paper and its Supplementary Information. The raw patient data are available from the authors, subject to Institutional Review Board approval. Source data for the figures are provided with this paper. The raw and analysed datasets generated during the study are available for research purposes from the corresponding authors on reasonable request. Source data are provided with this paper.
Code availability
The code for the custom macro generated for this study is provided in Supplementary Information.
References
Dong, X. Current strategies for brain drug delivery. Theranostics 8, 1481–1493 (2018).
Pardridge, W. M. Blood-brain barrier and delivery of protein and gene therapeutics to brain. Front. Aging Neurosci. 11, 373 (2019).
Sweeney, M. D., Zhao, Z., Montagne, A., Nelson, A. R. & Zlokovic, B. V. Blood-brain barrier: from physiology to disease and back. Physiol. Rev. 99, 21–78 (2019).
Varatharaj, A. & Galea, I. The blood-brain barrier in systemic inflammation. Brain Behav. Immun. 60, 1–12 (2017).
Nian, K., Harding, I. C., Herman, I. M. & Ebong, E. E. Blood-brain barrier damage in ischemic stroke and its regulation by endothelial mechanotransduction. Front. Physiol. 11, 605398 (2020).
Banks, W. A. et al. Transport of extracellular vesicles across the blood-brain barrier: brain pharmacokinetics and effects of inflammation. Int. J. Mol. Sci. 21, 4407 (2020).
Yuan, D. F. et al. Macrophage exosomes as natural nanocarriers for protein delivery to inflamed brain. Biomaterials 142, 1–12 (2017).
Haney, M. J. et al. Exosomes as drug delivery vehicles for Parkinson’s disease therapy. J. Control. Release 207, 18–30 (2015).
Man, S., Ubogu, E. E. & Ransohoff, R. M. Inflammatory cell migration into the central nervous system: a few new twists on an old tale. Brain Pathol. 17, 243–250 (2007).
Seguin, R., Biernacki, K., Rotondo, R. L., Prat, A. & Antel, J. P. Regulation and functional effects of monocyte migration across human brain-derived endothelial cells. J. Neuropathol. Exp. Neurol. 62, 412–419 (2003).
Erickson, M. A. & Banks, W. A. Age-associated changes in the immune system and blood-brain barrier functions. Int. J. Mol. Sci. 20, 1632 (2019).
Aslan, C. et al. Exosomes for mRNA delivery: a novel biotherapeutic strategy with hurdles and hope. BMC Biotechnol. 21, 20 (2021).
Li, M. et al. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Phil. Trans. R. Soc. Lond. B 369, 20130502 (2014).
Chevillet, J. R. et al. Quantitative and stoichiometric analysis of the microRNA content of exosomes. Proc. Natl Acad. Sci. USA 111, 14888–14893 (2014).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Luan, X. et al. Engineering exosomes as refined biological nanoplatforms for drug delivery. Acta Pharmacol. Sin. 38, 754–763 (2017).
Usman, W. M. et al. Efficient RNA drug delivery using red blood cell extracellular vesicles. Nat. Commun. 9, 2359 (2018).
Momen-Heravi, F., Bala, S., Bukong, T. & Szabo, G. Exosome-mediated delivery of functionally active miRNA-155 inhibitor to macrophages. Nanomedicine 10, 1517–1527 (2014).
Wang, J. H. et al. Anti-HER2 scFv-directed extracellular vesicle-mediated mRNA-based gene delivery inhibits growth of HER2-positive human breast tumor xenografts by prodrug activation. Mol. Cancer Ther. 17, 1133–1142 (2018).
Pastuzyn, E. D. et al. The neuronal gene arc encodes a repurposed retrotransposon gag protein that mediates intercellular RNA transfer. Cell 172, 275–288.e18 (2018).
Segel, M. et al. Mammalian retrovirus-like protein PEG10 packages its own mRNA and can be pseudotyped for mRNA delivery. Science 373, 882–889 (2021).
Ashley, J. et al. Retrovirus-like Gag Protein Arc1 binds RNA and traffics across synaptic boutons. Cell 172, 262–274.e11 (2018).
Comas-Garcia, M., Davis, S. R. & Rein, A. On the selective packaging of genomic RNA by HIV-1. Viruses 8, 246 (2016).
Dynes, J. L. & Steward, O. Arc mRNA docks precisely at the base of individual dendritic spines indicating the existence of a specialized microdomain for synapse-specific mRNA translation. J. Comp. Neurol. 520, 3105–3119 (2012).
Fila, M., Diaz, L., Szczepanska, J., Pawlowska, E. & Blasiak, J. mRNA trafficking in the nervous system: a key mechanism of the involvement of activity-regulated cytoskeleton-associated protein (Arc) in synaptic plasticity. Neural Plast. 2021, 3468795 (2021).
Paolantoni, C. et al. Arc 3’ UTR splicing leads to dual and antagonistic effects in fine-tuning arc expression upon BDNF signaling. Front. Mol. Neurosci. 11, 145 (2018).
Giorgi, C. et al. The EJC factor eIF4AIII modulates synaptic strength and neuronal protein expression. Cell 130, 179–191 (2007).
Booth, A. M. et al. Exosomes and HIV Gag bud from endosome-like domains of the T cell plasma membrane. J. Cell Biol. 172, 923–935 (2006).
Comas-Garcia, M. et al. Dissection of specific binding of HIV-1 Gag to the ‘packaging signal’ in viral RNA. Elife 6, e27055 (2017).
Brigham, B. S., Kitzrow, J. P., Reyes, J. C., Musier-Forsyth, K. & Munro, J. B. Intrinsic conformational dynamics of the HIV-1 genomic RNA 5’UTR. Proc. Natl Acad. Sci. USA 116, 10372–10381 (2019).
Blakemore, R. J. et al. Stability and conformation of the dimeric HIV-1 genomic RNA 5’UTR. Biophys. J. 120, 4874–4890 (2021).
Carlson, L. A., Bai, Y., Keane, S. C., Doudna, J. A. & Hurley, J. H. Reconstitution of selective HIV-1 RNA packaging in vitro by membrane-bound Gag assemblies. Elife 5, e14663 (2016).
Eriksen, M. S. et al. Arc self-association and formation of virus-like capsids are mediated by an N-terminal helical coil motif. FEBS J. 288, 2930–2955 (2021).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Rossler, K. et al. Expression of leucocyte adhesion molecules at the human blood-brain barrier (BBB). J. Neurosci. Res. 31, 365–374 (1992).
Granger, D. N. & Senchenkova, E. Inflammation and the Microcirculation (Morgan & Claypool Life Sciences, 2010).
Owens, T., Bechmann, I. & Engelhardt, B. Perivascular spaces and the two steps to neuroinflammation. J. Neuropathol. Exp. Neurol. 67, 1113–1121 (2008).
Zozulya, A. L. et al. Dendritic cell transmigration through brain microvessel endothelium is regulated by MIP-1alpha chemokine and matrix metalloproteinases. J. Immunol. 178, 520–529 (2007).
Larochelle, C., Alvarez, J. I. & Prat, A. How do immune cells overcome the blood-brain barrier in multiple sclerosis? FEBS Lett. 585, 3770–3780 (2011).
Acknowledgements
We thank S. Wilkens from SUNY Upstate Medical University for assistance with TEM imaging; R. Williams, T. Abratte and the BRC Imaging Facility at the Cornell Institute of Biotechnology (RRID:SCR_021741) for imaging experiments, with NIH S10OD025049 for the IVIS-spectrum optical imager in Cornell’s BRC Imaging Facility; P. Schweitzer and the BRC Genomics Facility (RRID:SCR_021727) at the Cornell Institute of Biotechnology for the sequencing experiments; L. Tesfa, J. Mahoney and the BRC Flow Cytometry Facility (RRID:SCR_021740) for flow cytometry data; T. Totman, E. Feldman, F. Burgus and the CARE at Cornell for service and advice on care of animals; R. Felt and the IACUC, Cornell for help composing and managing animal protocols; A. Recknagel for advice on imaging; Herbert Fountain for inspiring discussions about anti-aging therapies; and P. Miller for help in maintaining high-standard laboratory practices. S.J. acknowledges start-up support from Cornell University, including the Robert Langer ’70 Family and Friends Professorship, Cornell NEXT Nano Initiative and Cornell Engineering’s inaugural Sprout Awards.
Author information
Authors and Affiliations
Contributions
W. Gu, R.L. and S.J. conceptualized the work. W. Gu, S.L., K.L., S.C., N.E., Y.Y., M.C., Y.-W.C., W. Gao and T.S. acquired the data. W. Gu, S.L., S.C., Y.Y., Y.-W.C. and W. Gao contributed to data analyses and to the interpretation of the results. W. Gu, Z.Y., K.S.-G., R.H., P.M., C.W., C. Seo, A.G., C. Schaffer, N.N., R.C., Q.Y., M.W., R.L. and S.J. provided advice, support and supervision. W. Gu wrote the manuscript. W. Gu, C. Schaffer, N.N., R.C., M.W., R.L. and S.J. edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.J., W. Gu, S.L. and Z.Y. are authors of a patent application related to this work (PCT/US2022/027568) filed by Cornell University. All other authors declare no competing interests.
Peer review
Peer review information
Nature Biomedical Engineering thanks Koen Breyne, Takahiro Ochiya and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Optimization of CMDR dye staining for the quantification of total EV uptake.
To aid in quantifying EVs both in solution and within recipient cells, we employed plasma membrane staining using CMDR. We further refined the number of EVs introduced to recipient cells. The final parameters settled on a 1:5000 CMDR dye concentration and a 2X EV concentration, indicating EVs derived from 2 donor cells (over a span of 24-40 hours) were designated for a single recipient cell.
Extended Data Fig. 2 Density-related expression of the EV markers CD63 and CD81 in capsid+/ stabilizer+ eraEVs.
(a) Western blot analysis illustrating CD63 and Arc expression across various EV subpopulations, differentiated by density through density gradient centrifugation. The 70% sucrose cushion layer—representing the highest density—displayed a marked increase in both CD63 and Arc. Equal amounts of total proteins were loaded across all lanes. Displayed is one representative trial out of three independent experiments. (b) Immunomagnetic positive selection was conducted to isolate CD81+ EVs from filtered supernatant cell culture media laden with both control and engineered EVs. Following isolation, the CD81+ EVs were lysed to facilitate protein quantification and subsequent western blot analysis. (c) Utilizing the Pierce 660 nm Protein Assay, total proteins harvested from CD81+ EVs were quantified. Notably, there was a 2.3-fold upsurge in the protein count extracted from CD81 + /Arc+ engineered EVs in contrast to the capsid‒ control. (d) Western blotting of the CD81 + EV lysates affirmed the presence of Arc proteins. The absence of reducing agents in the sample buffer was deliberate since detection antibodies for CD9/CD63/CD81 often identify the disulfide bond associated with antigen epitopes. Under non-reducing conditions, substantial Arc oligomers are evident. Notably, despite comparable levels of monomer Arc, the quantity of Arc oligomers was significantly amplified in stabilizer+ EVs, underscoring the stabilization effect imparted by the A5U motif. Owing to our exclusion of an Arc knockout cell line, endogenous Arc was discernible in the no-transfection control set. Abbreviations: NT - no transfection control; AA - Arc/A5U-GFP; AG - Arc/GFP. (e-f) Quantitative breakdown depicting the distribution of Arc protein monomers and oligomers in each sample set, as determined through Western blot analysis. The figure presents one representative trial from three independent repeats. For uncropped western gels, refer to SD_ED_FIG2.
Supplementary information
Source data
Source Data for Extended Data Fig. 2
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gu, W., Luozhong, S., Cai, S. et al. Extracellular vesicles incorporating retrovirus-like capsids for the enhanced packaging and systemic delivery of mRNA into neurons. Nat. Biomed. Eng 8, 415–426 (2024). https://doi.org/10.1038/s41551-023-01150-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-023-01150-x